4.4: Dummy Variable Regression
- Page ID
- 33492
\[Y_{ij} = \mu + \mu_{i} + \epsilon_{ij}, \text{ fitted as } Y_{ij} = \beta_{0} + \beta_{Level \ 1} + \beta_{Level \ 2} + \ldots + \beta_{Level \ T-1} + \epsilon_{ij}\]
where \(\beta_{Level \ 1}, \beta_{Level \ 2}, \ldots, \beta_{Level \ T-1}\) are regression coefficients for \(T-1\) indicator-coded regression "dummy" variables that are correspond to the \(T-1\) categorical factor levels. The \(T^{th}\) factor level mean is given by the regression intercept \(\beta_{0}\).
The General Linear Model (GLM) applied to data with categorical predictors can be viewed from a regression modeling perspective as an ordinary multiple linear regression (MLR) with "dummy" coding, also known as indicator coding, for the categorical treatment levels. Typically, software performing the MLR will automatically include an intercept, which corresponds to the first column of the design matrix and is a column of \(1\)'s. This automatic inclusion of the intercept can lead to complications when interpreting the regression coefficients.
The SAS Mixed procedure, and also the GLM procedure which we may encounter later, use the "Dummy Variable Regression" model. For the \(Y\) data used in sections 4.2 and 4.3, the design matrix for this model can be entered into IML as:
/* Dummy Variable Regression Model */
x = {
1 1 0,
1 1 0,
1 0 1,
1 0 1,
1 0 0,
1 0 0};
Notice that in the above design matrix, there are only two indicator columns even though there are three treatment levels in the study. It is because, similar to the matrix below, if we were to have a design matrix with another indicator column representing the third treatment level, the resulting 4 columns would form a set of linearly dependent columns, a mathematical condition that will hinder the computation process any further as explained below. \[\begin{bmatrix} 1 & 1 & 0 & 0 \\ 1 & 1 & 0 & 0 \\ 1 & 0 & 1 & 0 \\ 1 & 0 & 1 & 0 \\ 1 & 0 & 0 & 1 \\ 1 & 0 & 0 & 1 \end{bmatrix}\]
The above matrix containing all 4 columns has the property that the sum of columns 2-4 will equal the first column representing the intercept. As a result, a mathematical condition called singularity is created and the matrix computations will not run. So one of the treatment levels is omitted from the coding in the design matrix above for IML and the eliminated level is called the ‘reference’ level. In SAS, typically, the treatment level with the highest label is defined as the reference level and so, in this study, it is treatment level 3.
Note that the parameter vector for the dummy variable regression model is \[\boldsymbol{\beta} = \begin{bmatrix} \mu_{1} \\ \mu_{2} \\ \mu_{3} \end{bmatrix}\].
Running IML, with the design matrix for the dummy variable regression model, we get the following output;
| Regression Coefficients | |
|---|---|
| Beta_0 | 5.5 |
| Beta_1 | -4 |
| Beta_2 | -2 |
The coefficient \(\beta_{0}\) is the mean for treatment level 3. The mean for treatment level 1 is then calculated from \(\hat{\beta}_{0} + \hat{\beta}_{1} = 1.5\). Likewise, the mean for treatment level 2 is calculated as \(\hat{\beta}_{0} + \hat{\beta}_{2} = 3.5\).
Notice that the \(F\) statistic calculated from this model is the same as that produced from the Cell Means model.
| ANOVA | ||||
|---|---|---|---|---|
| Treatment | df | SS | MS | F |
| 2 | 16 | 8 | 16 | |
| Error | 3 | 1.5 | 0.5 | |
| Total | 5 | 17.5 | ||
Using Technology
We can confirm our ANOVA table now by running the analysis in software such as Minitab.
- Steps in Minitab
-
First input the data:
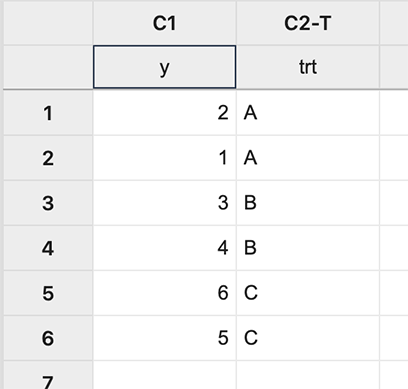
Figure \(\PageIndex{a1}\): Inputting data. In Minitab, different coding options allow the choice of the design matrix which can be done as follows:
Stat > ANOVA > General Linear Model > Fit General Linear Model and place the variables in the appropriate boxes:
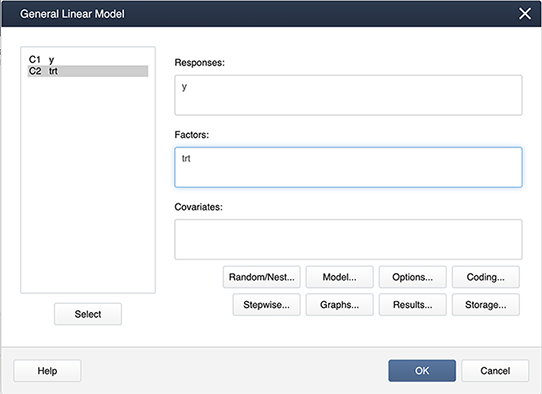
Figure \(\PageIndex{a2}\): Placing variables in the General Linear Model pop-up window. Then select Coding… and choose the (1,0) coding as shown below:
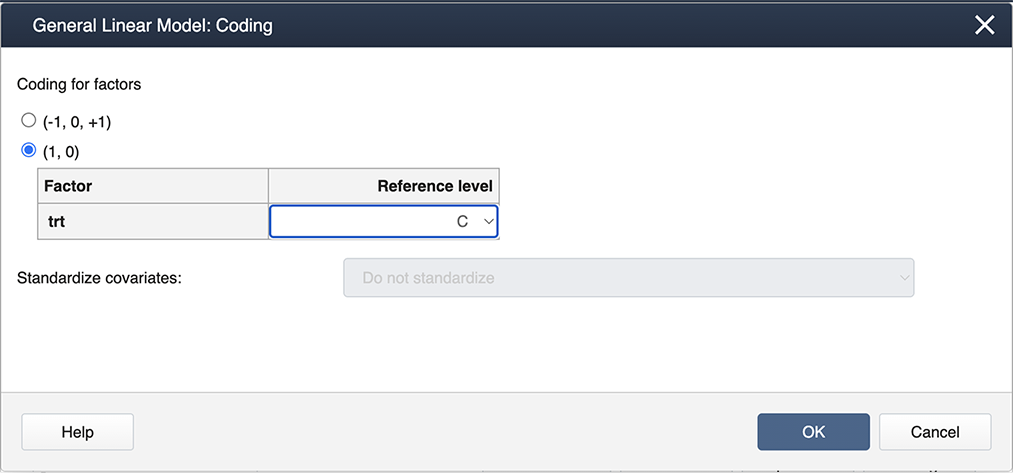
Figure \(\PageIndex{a3}\): Selecting options in the General Linear Model: Coding window. Select OK to exit the nested windows. This produces the regular ANOVA output:
Analysis of Variance
Source DF Adj SS Adj MS F-Value P-Value trt 2 16.000 8.0000 16.00 0.025 Error 3 1.500 0.5000 Total 5 17.500 And also the Regression Equation:
Regression Equation
y = 5.500 - 4.000 trt_level1 - 2.000 trt_level2 + 0.0 trt_level3
- Steps in SAS
-
In SAS, the default coding is indicator coding, so when you specify the option
model y=trt / solution;
you get the regression coefficients:
Solution for Fixed Effects Effect trt Estimate Standard Error DF t Value Pr > |t| Intercept 5.5000 0.5000 3 11.00 0.0016 trt level1 -4.0000 0.7071 3 -5.66 0.0109 trt level2 -2.0000 0.7071 3 -2.83 0.0663 trt level3 0 And the same ANOVA table:
Type 3 Analysis of Variance Source DF Sum of Squares Mean Square Expected Mean Square Error Term Error DF F Value Pr > F trt 2 16.000000 8.000000 Var(Residual)+Q(trt) MS(Residual) 3 16.00 0.0251 Residual 3 1.500000 0.500000 Var(Residual) The Intermediate calculations for this model are:
xprimex 6 2 2 2 2 0 2 0 2 check 1 -2.22E-16 0 3.331E-16 1 0 0 0 1 xprimey 21 3 7 SumY2 89.5 CF 73.5 xprimexinv 0.5 -0.5 -0.5 -0.5 1 0.5 -0.5 0.5 1
- Steps in R
-
1. Define response variable and design matrix
y<-matrix(c(2,1,3,4,6,5), ncol=1) x = matrix(c(1,1,0,1,1,0,1,0,1,1,0,1,1,0,0,1,0,0),ncol=3,nrow=6,byrow=TRUE)
2. Regression coefficients
beta<-solve(t(x)%*%x)%*%(t(x)%*%y) # beta # [,1] # [1,] 5.5 # [2,] -4.0 # [3,] -2.03. Calculate the entries of the ANOVA Table
n<-nrow(y) p<-ncol(x) J<-matrix(1,n,n) ss_tot = (t(y)%*%y) - (1/n)*(t(y)%*%J)%*%y #17.5 ss_trt = t(beta)%*%(t(x)%*%y) - (1/n)*(t(y)%*%J)%*%y #16 ss_error = ss_tot - ss_trt #1.5 total_df=n-1 #5 trt_df=p-1 #2 error_df=n-p #3 MS_trt = ss_trt/(p-1) #8 MS_error = ss_error / error_df #0.5 F=MS_trt/MS_error #16
4. Creating the ANOVA table
ANOVA <- data.frame( c ("","Treatment","Error", "Total"), c("DF", trt_df,error_df,total_df), c("SS", ss_trt, ss_error, ss_tot), c("MS", MS_trt, MS_error, ""), c("F",F,"",""), stringsAsFactors = FALSE) names(ANOVA) <- c(" ", " ", " ","","")5. Print the ANOVA table
print(ANOVA) # 1 DF SS MS F # 2 Treatment 2 16 8 16 # 3 Error 3 1.5 0.5 # 4 Total 5 17.56. Intermediates in the matrix computations
xprimex<-t(x)%*%x # xprimex # [,1] [,2] [,3] # [1,] 6 2 2 # [2,] 2 2 0 # [3,] 2 0 2 xprimey<-t(x)%*%y # xprimey # [,1] # [1,] 21 # [2,] 3 # [3,] 7 xprimexinv<-solve(t(x)%*%x) # xprimexinv # [,1] [,2] [,3] # [1,] 0.5 -0.5 -0.5 # [2,] -0.5 1.0 0.5 # [3,] -0.5 0.5 1.0 check<-xprimexinv%*%xprimex # check # [,1] [,2] [,3] # [1,] 1.000000e+00 0.000000e+00 0 # [2,] -1.110223e-16 1.000000e+00 0 # [3,] 0.000000e+00 -1.110223e-16 1 SumY2<-t(beta)%*%(t(x)%*%y) # 89.5 CF<-(1/n)*(t(y)%*%J)%*%y # 73.5
7. Regression Equation and ANOVA table
trt_level1<-x[,2] trt_level2<-x[,3] model<-lm(y~trt_level1+trt_level2)
8. With the command summary(model) we can get the following output:
Call: lm(formula = y ~ trt_level1 + trt_level2) Residuals: 1 2 3 4 5 6 0.5 -0.5 -0.5 0.5 0.5 -0.5 Coefficients: Estimate Std. Error t value Pr(>|t|) (Intercept) 5.5000 0.5000 11.000 0.00161 ** trt_level1 -4.0000 0.7071 -5.657 0.01094 * trt_level2 -2.0000 0.7071 -2.828 0.06628 . --- Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1 Residual standard error: 0.7071 on 3 degrees of freedom Multiple R-squared: 0.9143, Adjusted R-squared: 0.8571 F-statistic: 16 on 2 and 3 DF, p-value: 0.02509
From the output, we can see the estimates for the coefficients are b0=5.5, b1=-4, b2=-2 and the F-statistic is 16 with a p-value of 0.02509.
By using the estimates we can write the regression equation:
y=5.5-4 trt_level1-2 trt_level2+0 trt_level3
-
9. With the command anova(model) we can get the following output
Analysis of Variance Table Response: y Df Sum Sq Mean Sq F value Pr(>F) trt_level1 1 12.0 12.0 24 0.01628 * trt_level2 1 4.0 4.0 8 0.06628 . Residuals 3 1.5 0.5 --- Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Note: R is giving the sequential sum of squares in the ANOVA table.


