3.9: Answers to exercises
- Page ID
- 3914
Answers to the barplot coloring question:
Code \(\PageIndex{1}\) (R):
or
Code \(\PageIndex{2}\) (R):
(Please try these commands yourself. The second answer is preferable because it will work even in cases when factor has more than two levels.)
Answers to the barplot counts question. To see frequencies, from highest to smallest, run:
Code \(\PageIndex{3}\) (R):
or
Code \(\PageIndex{4}\) (R):
Answer to flowering heads question. First, we need to load the file into R. With url.show() or simply by examining the file in the browser window, we reveal that file has multiple columns divided with wide spaces (likely Tab symbols) and that the first column contains species names with spaces. Therefore, header and separator should be defined explicitly:
Code \(\PageIndex{5}\) (R):
Next step is always to check the structure of new object:
Code \(\PageIndex{6}\) (R):
Two columns (including species) are factors, others are numerical (integer or not). The resulted object is a data frame.
Next is to select our species and remove unused levels:
Code \(\PageIndex{7}\) (R):
To select tree species in one command, we used logical expression made with %in% operator (please see how it works with ?"%in%" command).
Removal of redundant levels will help to use species names for scatterplot:
Code \(\PageIndex{8}\) (R):
Please make this plot yourself. The key is to use SPECIES factor as number, with as.numeric() command. Function with() allows to ignore cc$ and therefore saves typing.
However, there is one big problem which at first is not easy to recognize: in many places, points overlay each other and therefore amount of visible data points is much less then in the data file. What is worse, we cannot say if first and third species are well or not well segregated because we do not see how many data values are located on the “border” between them. This scatterplot problem is well known and there are workarounds:
Code \(\PageIndex{9}\) (R):
Please run this code yourself. Function jitter() adds random noise to variables and shits points allowing to see what is below. However, it is still hard to understand the amount of overplotted values.
There are also:
Code \(\PageIndex{10}\) (R):
(Try these variants yourself. When you run the first line of code, you will see sunflower plot, developed exactly to such “overplotting cases”. It reflects how many points are overlaid. However, it is not easy to make sunflowerplot() show overplotting separately for each species. The other approach, smoothScatter() suffers from the same problem\(^{[1]}\).)
To overcome this, we developed PPoints() function (Figure \(\PageIndex{1}\)):
Code \(\PageIndex{11}\) (R):
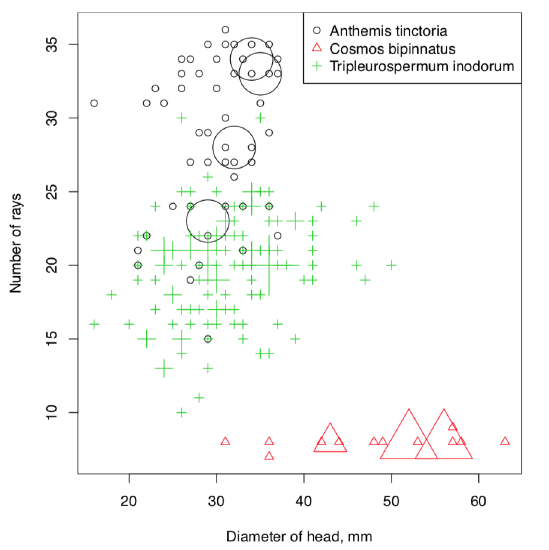
Finally, the answer. As one might see, garden cosmos is really separate from two other species which in turn could be distinguished with some certainty, mostly because number of rays in the yellow chamomile is more than 20. This approach is possible to improve. “Data mining” chapter tells how to do that.
Answer to the matrix question. While creating matrix ma, we defined byrow=TRUE, i.e. indicated that elements should be joined into a matrix row by row. In case of byrow=FALSE (default) we would have obtained the matrix identical to mb:
Code \(\PageIndex{12}\) (R):
Answer to the sorting exercise. To work with columns, we have to use square brackets with a comma and place commands to the right:
Code \(\PageIndex{13}\) (R):
Please note that we cannot just type order() after the comma. This command returns the new order of columns, thus we gave it our column names (names() returns column names for a given data frame). By the way, sort() would have worked here too, since we only needed to rearrange a single vector.
References
1. There is also hexbin package which used hexagonal shapes and color shading.

