12.1: The χ2 Goodness-of-fit Test
- Page ID
- 4014
The χ2 goodness-of-fit test is one of the oldest hypothesis tests around: it was invented by Karl Pearson around the turn of the century (Pearson 1900), with some corrections made later by Sir Ronald Fisher (Fisher 1922a). To introduce the statistical problem that it addresses, let’s start with some psychology…
cards data
Over the years, there have been a lot of studies showing that humans have a lot of difficulties in simulating randomness. Try as we might to “act” random, we think in terms of patterns and structure, and so when asked to “do something at random”, what people actually do is anything but random. As a consequence, the study of human randomness (or non-randomness, as the case may be) opens up a lot of deep psychological questions about how we think about the world. With this in mind, let’s consider a very simple study. Suppose I asked people to imagine a shuffled deck of cards, and mentally pick one card from this imaginary deck “at random”. After they’ve chosen one card, I ask them to mentally select a second one. For both choices, what we’re going to look at is the suit (hearts, clubs, spades or diamonds) that people chose. After asking, say, N=200 people to do this, I’d like to look at the data and figure out whether or not the cards that people pretended to select were really random. The data are contained in the randomness.Rdata file, which contains a single data frame called cards. Let’s take a look:
library( lsr )
load( "./rbook-master/data/randomness.Rdata" )
str(cards)## 'data.frame': 200 obs. of 3 variables:
## $ id : Factor w/ 200 levels "subj1","subj10",..: 1 112 124 135 146 157 168 179 190 2 ...
## $ choice_1: Factor w/ 4 levels "clubs","diamonds",..: 4 2 3 4 3 1 3 2 4 2 ...
## $ choice_2: Factor w/ 4 levels "clubs","diamonds",..: 1 1 1 1 4 3 2 1 1 4 ...As you can see, the cards data frame contains three variables, an id variable that assigns a unique identifier to each participant, and the two variables choice_1 and choice_2 that indicate the card suits that people chose. Here’s the first few entries in the data frame:
head( cards )## id choice_1 choice_2
## 1 subj1 spades clubs
## 2 subj2 diamonds clubs
## 3 subj3 hearts clubs
## 4 subj4 spades clubs
## 5 subj5 hearts spades
## 6 subj6 clubs heartsFor the moment, let’s just focus on the first choice that people made. We’ll use the table() function to count the number of times that we observed people choosing each suit. I’ll save the table to a variable called observed, for reasons that will become clear very soon:
observed <- table( cards$choice_1 )
observed##
## clubs diamonds hearts spades
## 35 51 64 50That little frequency table is quite helpful. Looking at it, there’s a bit of a hint that people might be more likely to select hearts than clubs, but it’s not completely obvious just from looking at it whether that’s really true, or if this is just due to chance. So we’ll probably have to do some kind of statistical analysis to find out, which is what I’m going to talk about in the next section.
Excellent. From this point on, we’ll treat this table as the data that we’re looking to analyse. However, since I’m going to have to talk about this data in mathematical terms (sorry!) it might be a good idea to be clear about what the notation is. In R, if I wanted to pull out the number of people that selected diamonds, I could do it by name by typing observed["diamonds"] but, since "diamonds" is second element of the observed vector, it’s equally effective to refer to it as observed[2]. The mathematical notation for this is pretty similar, except that we shorten the human-readable word “observed” to the letter O, and we use subscripts rather than brackets: so the second observation in our table is written as observed[2] in R, and is written as O2 in maths. The relationship between the English descriptions, the R commands, and the mathematical symbols are illustrated below:
| label | index i | math. symbol | R command | the value |
|---|---|---|---|---|
| clubs ♣ | 1 | O1 | observed[1] |
35 |
| diamonds ♢ | 2 | O2 | observed[2] |
51 |
| hearts ♡ | 3 | O3 | observed[3] |
64 |
| spades ♠ | 4 | O4 | observed[4] |
50 |
Hopefully that’s pretty clear. It’s also worth nothing that mathematicians prefer to talk about things in general rather than specific things, so you’ll also see the notation Oi, which refers to the number of observations that fall within the i-th category (where i could be 1, 2, 3 or 4). Finally, if we want to refer to the set of all observed frequencies, statisticians group all of observed values into a vector, which I’ll refer to as O.
O=(O1,O2,O3,O4)
Again, there’s nothing new or interesting here: it’s just notation. If I say that O = (35,51,64,50) all I’m doing is describing the table of observed frequencies (i.e., observed), but I’m referring to it using mathematical notation, rather than by referring to an R variable.
null hypothesis and the alternative hypothesis
As the last section indicated, our research hypothesis is that “people don’t choose cards randomly”. What we’re going to want to do now is translate this into some statistical hypotheses, and construct a statistical test of those hypotheses. The test that I’m going to describe to you is Pearson’s χ2 goodness of fit test, and as is so often the case, we have to begin by carefully constructing our null hypothesis. In this case, it’s pretty easy. First, let’s state the null hypothesis in words:
| H0 |
|---|
| All four suits are chosen with equal probability |
Now, because this is statistics, we have to be able to say the same thing in a mathematical way. To do this, let’s use the notation Pj to refer to the true probability that the j-th suit is chosen. If the null hypothesis is true, then each of the four suits has a 25% chance of being selected: in other words, our null hypothesis claims that P1=.25, P2=.25, P3=.25 and finally that P4=.25. However, in the same way that we can group our observed frequencies into a vector O that summarises the entire data set, we can use P to refer to the probabilities that correspond to our null hypothesis. So if I let the vector P=(P1,P2,P3,P4) refer to the collection of probabilities that describe our null hypothesis, then we have
H0:P=(.25,.25,.25,.25)
In this particular instance, our null hypothesis corresponds to a vector of probabilities P in which all of the probabilities are equal to one another. But this doesn’t have to be the case. For instance, if the experimental task was for people to imagine they were drawing from a deck that had twice as many clubs as any other suit, then the null hypothesis would correspond to something like P=(.4,.2,.2,.2). As long as the probabilities are all positive numbers, and they all sum to 1, them it’s a perfectly legitimate choice for the null hypothesis. However, the most common use of the goodness of fit test is to test a null hypothesis that all of the categories are equally likely, so we’ll stick to that for our example.
What about our alternative hypothesis, H1? All we’re really interested in is demonstrating that the probabilities involved aren’t all identical (that is, people’s choices weren’t completely random). As a consequence, the “human friendly” versions of our hypotheses look like this:
| H0 | H1 |
|---|---|
| All four suits are chosen with equal probability and the “mathematician friendly” version is |
At least one of the suit-choice probabilities isn’t .25 |
| H0 | H1 |
|---|---|
| P=(.25,.25,.25,.25) | P≠(.25,.25,.25,.25) |
Conveniently, the mathematical version of the hypotheses looks quite similar to an R command defining a vector. So maybe what I should do is store the P vector in R as well, since we’re almost certainly going to need it later. And because I’m so imaginative, I’ll call this R vector probabilities,
probabilities <- c(clubs = .25, diamonds = .25, hearts = .25, spades = .25)
probabilities## clubs diamonds hearts spades
## 0.25 0.25 0.25 0.25“goodness of fit” test statistic
At this point, we have our observed frequencies O and a collection of probabilities P corresponding the null hypothesis that we want to test. We’ve stored these in R as the corresponding variables observed and probabilities. What we now want to do is construct a test of the null hypothesis. As always, if we want to test H0 against H1, we’re going to need a test statistic. The basic trick that a goodness of fit test uses is to construct a test statistic that measures how “close” the data are to the null hypothesis. If the data don’t resemble what you’d “expect” to see if the null hypothesis were true, then it probably isn’t true. Okay, if the null hypothesis were true, what would we expect to see? Or, to use the correct terminology, what are the expected frequencies. There are N=200 observations, and (if the null is true) the probability of any one of them choosing a heart is P3=.25, so I guess we’re expecting 200×.25=50 hearts, right? Or, more specifically, if we let Ei refer to “the number of category i responses that we’re expecting if the null is true”, then
Ei=N×Pi
This is pretty easy to calculate in R:
N <- 200 # sample size
expected <- N * probabilities # expected frequencies
expected## clubs diamonds hearts spades
## 50 50 50 50None of which is very surprising: if there are 200 observation that can fall into four categories, and we think that all four categories are equally likely, then on average we’d expect to see 50 observations in each category, right?
Now, how do we translate this into a test statistic? Clearly, what we want to do is compare the expected number of observations in each category (Ei) with the observed number of observations in that category (Oi). And on the basis of this comparison, we ought to be able to come up with a good test statistic. To start with, let’s calculate the difference between what the null hypothesis expected us to find and what we actually did find. That is, we calculate the “observed minus expected” difference score, Oi−Ei. This is illustrated in the following table.
| ♣ | ♢ | ♡ | ♠ | ||
| expected frequency | Ei | 50 | 50 | 50 | 50 |
| observed frequency | Oi | 35 | 51 | 64 | 50 |
| difference score | Oi−Ei | -15 | 1 | 14 | 0 |
The same calculations can be done in R, using our expected and observed variables:
observed - expected ##
## clubs diamonds hearts spades
## -15 1 14 0Regardless of whether we do the calculations by hand or whether we do them in R, it’s clear that people chose more hearts and fewer clubs than the null hypothesis predicted. However, a moment’s thought suggests that these raw differences aren’t quite what we’re looking for. Intuitively, it feels like it’s just as bad when the null hypothesis predicts too few observations (which is what happened with hearts) as it is when it predicts too many (which is what happened with clubs). So it’s a bit weird that we have a negative number for clubs and a positive number for heards. One easy way to fix this is to square everything, so that we now calculate the squared differences, (Ei−Oi)2. As before, we could do this by hand, but it’s easier to do it in R…
(observed - expected)^2##
## clubs diamonds hearts spades
## 225 1 196 0Now we’re making progress. What we’ve got now is a collection of numbers that are big whenever the null hypothesis makes a bad prediction (clubs and hearts), but are small whenever it makes a good one (diamonds and spades). Next, for some technical reasons that I’ll explain in a moment, let’s also divide all these numbers by the expected frequency Ei, so we’re actually calculating \(\dfrac{\left(E_{i}-O_{i}\right)^{2}}{E_{i}}\). Since Ei=50 for all categories in our example, it’s not a very interesting calculation, but let’s do it anyway. The R command becomes:
(observed - expected)^2 / expected##
## clubs diamonds hearts spades
## 4.50 0.02 3.92 0.00In effect, what we’ve got here are four different “error” scores, each one telling us how big a “mistake” the null hypothesis made when we tried to use it to predict our observed frequencies. So, in order to convert this into a useful test statistic, one thing we could do is just add these numbers up. The result is called the goodness of fit statistic, conventionally referred to either as X2 or GOF. We can calculate it using this command in R
sum( (observed - expected)^2 / expected )## [1] 8.44The formula for this statistic looks remarkably similar to the R command. If we let k refer to the total number of categories (i.e., k=4 for our cards data), then the X2 statistic is given by:
\(X^{2}=\sum_{i=1}^{k} \dfrac{\left(O_{i}-E_{i}\right)^{2}}{E_{i}}\)
Intuitively, it’s clear that if X2 is small, then the observed data Oi are very close to what the null hypothesis predicted Ei, so we’re going to need a large X2 statistic in order to reject the null. As we’ve seen from our calculations, in our cards data set we’ve got a value of X2=8.44. So now the question becomes, is this a big enough value to reject the null?
sampling distribution of the GOF statistic (advanced)
To determine whether or not a particular value of X2 is large enough to justify rejecting the null hypothesis, we’re going to need to figure out what the sampling distribution for X2 would be if the null hypothesis were true. So that’s what I’m going to do in this section. I’ll show you in a fair amount of detail how this sampling distribution is constructed, and then – in the next section – use it to build up a hypothesis test. If you want to cut to the chase and are willing to take it on faith that the sampling distribution is a chi-squared (χ2) distribution with k−1 degrees of freedom, you can skip the rest of this section. However, if you want to understand why the goodness of fit test works the way it does, read on…
Okay, let’s suppose that the null hypothesis is actually true. If so, then the true probability that an observation falls in the i-th category is Pi – after all, that’s pretty much the definition of our null hypothesis. Let’s think about what this actually means. If you think about it, this is kind of like saying that “nature” makes the decision about whether or not the observation ends up in category i by flipping a weighted coin (i.e., one where the probability of getting a head is Pj). And therefore, we can think of our observed frequency Oi by imagining that nature flipped N of these coins (one for each observation in the data set)… and exactly Oi of them came up heads. Obviously, this is a pretty weird way to think about the experiment. But what it does (I hope) is remind you that we’ve actually seen this scenario before. It’s exactly the same set up that gave rise to the binomial distribution in Section 9.4. In other words, if the null hypothesis is true, then it follows that our observed frequencies were generated by sampling from a binomial distribution:
Oi∼Binomial(Pi,N)
Now, if you remember from our discussion of the central limit theorem (Section 10.3.3), the binomial distribution starts to look pretty much identical to the normal distribution, especially when N is large and when Pi isn’t too close to 0 or 1. In other words as long as N×Pi is large enough – or, to put it another way, when the expected frequency Ei is large enough – the theoretical distribution of Oi is approximately normal. Better yet, if Oi is normally distributed, then so is (Oi−Ei)/\(\sqrt{E_{i}}\) … since Ei is a fixed value, subtracting off Ei and dividing by \(\sqrt{E_{i}}\) changes the mean and standard deviation of the normal distribution; but that’s all it does. Okay, so now let’s have a look at what our goodness of fit statistic actually is. What we’re doing is taking a bunch of things that are normally-distributed, squaring them, and adding them up. Wait. We’ve seen that before too! As we discussed in Section 9.6, when you take a bunch of things that have a standard normal distribution (i.e., mean 0 and standard deviation 1), square them, then add them up, then the resulting quantity has a chi-square distribution. So now we know that the null hypothesis predicts that the sampling distribution of the goodness of fit statistic is a chi-square distribution. Cool.
There’s one last detail to talk about, namely the degrees of freedom. If you remember back to Section 9.6, I said that if the number of things you’re adding up is k, then the degrees of freedom for the resulting chi-square distribution is k. Yet, what I said at the start of this section is that the actual degrees of freedom for the chi-square goodness of fit test is k−1. What’s up with that? The answer here is that what we’re supposed to be looking at is the number of genuinely independent things that are getting added together. And, as I’ll go on to talk about in the next section, even though there’s k things that we’re adding, only k−1 of them are truly independent; and so the degrees of freedom is actually only k−1. That’s the topic of the next section.170
Degrees of freedom
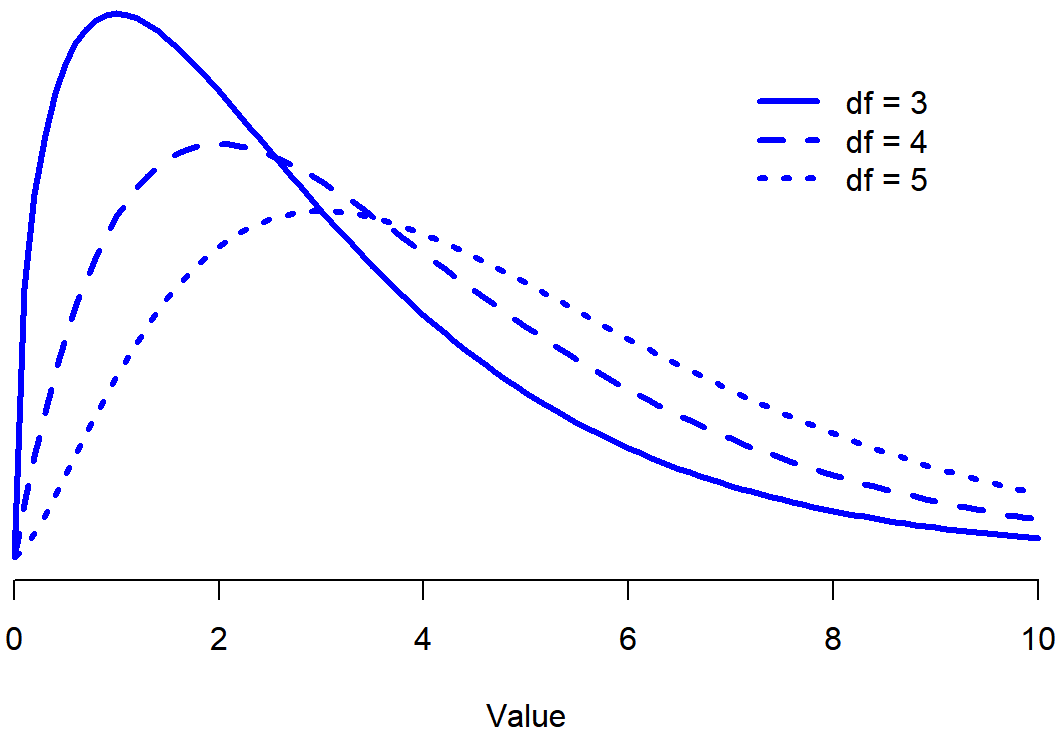
When I introduced the chi-square distribution in Section 9.6, I was a bit vague about what “degrees of freedom” actually means. Obviously, it matters: looking Figure 12.1 you can see that if we change the degrees of freedom, then the chi-square distribution changes shape quite substantially. But what exactly is it? Again, when I introduced the distribution and explained its relationship to the normal distribution, I did offer an answer… it’s the number of “normally distributed variables” that I’m squaring and adding together. But, for most people, that’s kind of abstract, and not entirely helpful. What we really need to do is try to understand degrees of freedom in terms of our data. So here goes.
The basic idea behind degrees of freedom is quite simple: you calculate it by counting up the number of distinct “quantities” that are used to describe your data; and then subtracting off all of the “constraints” that those data must satisfy.171 This is a bit vague, so let’s use our cards data as a concrete example. We describe out data using four numbers, O1, O2, O3 and O4 corresponding to the observed frequencies of the four different categories (hearts, clubs, diamonds, spades). These four numbers are the random outcomes of our experiment. But, my experiment actually has a fixed constraint built into it: the sample size N.172 That is, if we know how many people chose hearts, how many chose diamonds and how many chose clubs; then we’d be able to figure out exactly how many chose spades. In other words, although our data are described using four numbers, they only actually correspond to 4−1=3 degrees of freedom. A slightly different way of thinking about it is to notice that there are four probabilities that we’re interested in (again, corresponding to the four different categories), but these probabilities must sum to one, which imposes a constraint. Therefore, the degrees of freedom is 4−1=3. Regardless of whether you want to think about it in terms of the observed frequencies or in terms of the probabilities, the answer is the same. In general, when running the chi-square goodness of fit test for an experiment involving k groups, then the degrees of freedom will be k−1.
Testing the null hypothesis
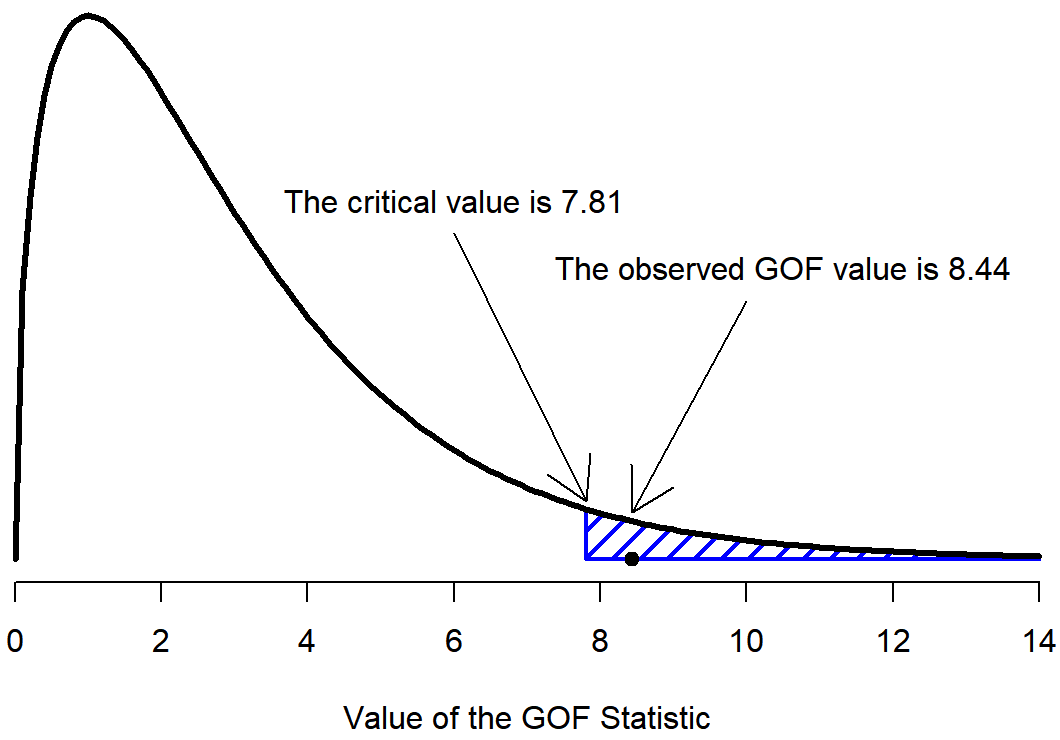
The final step in the process of constructing our hypothesis test is to figure out what the rejection region is. That is, what values of X2 would lead is to reject the null hypothesis. As we saw earlier, large values of X2 imply that the null hypothesis has done a poor job of predicting the data from our experiment, whereas small values of X2 imply that it’s actually done pretty well. Therefore, a pretty sensible strategy would be to say there is some critical value, such that if X2 is bigger than the critical value we reject the null; but if X2 is smaller than this value we retain the null. In other words, to use the language we introduced in Chapter @ref(hypothesistesting the chi-squared goodness of fit test is always a one-sided test. Right, so all we have to do is figure out what this critical value is. And it’s pretty straightforward. If we want our test to have significance level of α=.05 (that is, we are willing to tolerate a Type I error rate of 5%), then we have to choose our critical value so that there is only a 5% chance that X2 could get to be that big if the null hypothesis is true. That is to say, we want the 95th percentile of the sampling distribution. This is illustrated in Figure 12.2.
Ah, but – I hear you ask – how do I calculate the 95th percentile of a chi-squared distribution with k−1 degrees of freedom? If only R had some function, called… oh, I don’t know, qchisq() … that would let you calculate this percentile (see Chapter 9 if you’ve forgotten). Like this…
qchisq( p = .95, df = 3 )## [1] 7.814728So if our X2 statistic is bigger than 7.81 or so, then we can reject the null hypothesis. Since we actually calculated that before (i.e., X2=8.44) we can reject the null. If we want an exact p-value, we can calculate it using the pchisq() function:
pchisq( q = 8.44, df = 3, lower.tail = FALSE )## [1] 0.03774185This is hopefully pretty straightforward, as long as you recall that the “p” form of the probability distribution functions in R always calculates the probability of getting a value of less than the value you entered (in this case 8.44). We want the opposite: the probability of getting a value of 8.44 or more. That’s why I told R to use the upper tail, not the lower tail. That said, it’s usually easier to calculate the p-value this way:
1-pchisq( q = 8.44, df = 3 )## [1] 0.03774185So, in this case we would reject the null hypothesis, since p<.05. And that’s it, basically. You now know “Pearson’s χ2 test for the goodness of fit”. Lucky you.
Doing the test in R
Gosh darn it. Although we did manage to do everything in R as we were going through that little example, it does rather feel as if we’re typing too many things into the magic computing box. And I hate typing. Not surprisingly, R provides a function that will do all of these calculations for you. In fact, there are several different ways of doing it. The one that most people use is the chisq.test() function, which comes with every installation of R. I’ll show you how to use the chisq.test() function later on (in Section @ref(chisq.test), but to start out with I’m going to show you the goodnessOfFitTest() function in the lsr package, because it produces output that I think is easier for beginners to understand. It’s pretty straightforward: our raw data are stored in the variable cards$choice_1, right? If you want to test the null hypothesis that all four suits are equally likely, then (assuming you have the lsr package loaded) all you have to do is type this:
goodnessOfFitTest( cards$choice_1 ) ##
## Chi-square test against specified probabilities
##
## Data variable: cards$choice_1
##
## Hypotheses:
## null: true probabilities are as specified
## alternative: true probabilities differ from those specified
##
## Descriptives:
## observed freq. expected freq. specified prob.
## clubs 35 50 0.25
## diamonds 51 50 0.25
## hearts 64 50 0.25
## spades 50 50 0.25
##
## Test results:
## X-squared statistic: 8.44
## degrees of freedom: 3
## p-value: 0.038R then runs the test, and prints several lines of text. I’ll go through the output line by line, so that you can make sure that you understand what you’re looking at. The first two lines are just telling you things you already know:
Chi-square test against specified probabilities
Data variable: cards$choice_1 The first line tells us what kind of hypothesis test we ran, and the second line tells us the name of the variable that we ran it on. After that comes a statement of what the null and alternative hypotheses are:
Hypotheses:
null: true probabilities are as specified
alternative: true probabilities differ from those specified
For a beginner, it’s kind of handy to have this as part of the output: it’s a nice reminder of what your null and alternative hypotheses are. Don’t get used to seeing this though. The vast majority of hypothesis tests in R aren’t so kind to novices. Most R functions are written on the assumption that you already understand the statistical tool that you’re using, so they don’t bother to include an explicit statement of the null and alternative hypothesis. The only reason that goodnessOfFitTest() actually does give you this is that I wrote it with novices in mind.
The next part of the output shows you the comparison between the observed frequencies and the expected frequencies:
Descriptives:
observed freq. expected freq. specified prob.
clubs 35 50 0.25
diamonds 51 50 0.25
hearts 64 50 0.25
spades 50 50 0.25The first column shows what the observed frequencies were, the second column shows the expected frequencies according to the null hypothesis, and the third column shows you what the probabilities actually were according to the null. For novice users, I think this is helpful: you can look at this part of the output and check that it makes sense: if it doesn’t you might have typed something incorrecrtly.
The last part of the output is the “important” stuff: it’s the result of the hypothesis test itself. There are three key numbers that need to be reported: the value of the X2 statistic, the degrees of freedom, and the p-value:
Test results:
X-squared statistic: 8.44
degrees of freedom: 3
p-value: 0.038 Notice that these are the same numbers that we came up with when doing the calculations the long way.
Specifying a different null hypothesis
At this point you might be wondering what to do if you want to run a goodness of fit test, but your null hypothesis is not that all categories are equally likely. For instance, let’s suppose that someone had made the theoretical prediction that people should choose red cards 60% of the time, and black cards 40% of the time (I’ve no idea why you’d predict that), but had no other preferences. If that were the case, the null hypothesis would be to expect 30% of the choices to be hearts, 30% to be diamonds, 20% to be spades and 20% to be clubs. This seems like a silly theory to me, and it’s pretty easy to test it using our data. All we need to do is specify the probabilities associated with the null hypothesis. We create a vector like this:
nullProbs <- c(clubs = .2, diamonds = .3, hearts = .3, spades = .2)
nullProbs## clubs diamonds hearts spades
## 0.2 0.3 0.3 0.2Now that we have an explicitly specified null hypothesis, we include it in our command. This time round I’ll use the argument names properly. The data variable corresponds to the argument x, and the probabilities according to the null hypothesis correspond to the argument p. So our command is:
goodnessOfFitTest( x = cards$choice_1, p = nullProbs )
##
## Chi-square test against specified probabilities
##
## Data variable: cards$choice_1
##
## Hypotheses:
## null: true probabilities are as specified
## alternative: true probabilities differ from those specified
##
## Descriptives:
## observed freq. expected freq. specified prob.
## clubs 35 40 0.2
## diamonds 51 60 0.3
## hearts 64 60 0.3
## spades 50 40 0.2
##
## Test results:
## X-squared statistic: 4.742
## degrees of freedom: 3
## p-value: 0.192As you can see the null hypothesis and the expected frequencies are different to what they were last time. As a consequence our X2 test statistic is different, and our p-value is different too. Annoyingly, the p-value is .192, so we can’t reject the null hypothesis. Sadly, despite the fact that the null hypothesis corresponds to a very silly theory, these data don’t provide enough evidence against it.
report the results of the test
So now you know how the test works, and you know how to do the test using a wonderful magic computing box. The next thing you need to know is how to write up the results. After all, there’s no point in designing and running an experiment and then analysing the data if you don’t tell anyone about it! So let’s now talk about what you need to do when reporting your analysis. Let’s stick with our card-suits example. If I wanted to write this result up for a paper or something, the conventional way to report this would be to write something like this:
Of the 200 participants in the experiment, 64 selected hearts for their first choice, 51 selected diamonds, 50 selected spades, and 35 selected clubs. A chi-square goodness of fit test was conducted to test whether the choice probabilities were identical for all four suits. The results were significant (χ2(3)=8.44,p<.05), suggesting that people did not select suits purely at random.
This is pretty straightforward, and hopefully it seems pretty unremarkable. That said, there’s a few things that you should note about this description:
- The statistical test is preceded by the descriptive statistics. That is, I told the reader something about what the data look like before going on to do the test. In general, this is good practice: always remember that your reader doesn’t know your data anywhere near as well as you do. So unless you describe it to them properly, the statistical tests won’t make any sense to them, and they’ll get frustrated and cry.
- The description tells you what the null hypothesis being tested is. To be honest, writers don’t always do this, but it’s often a good idea in those situations where some ambiguity exists; or when you can’t rely on your readership being intimately familiar with the statistical tools that you’re using. Quite often the reader might not know (or remember) all the details of the test that your using, so it’s a kind of politeness to “remind” them! As far as the goodness of fit test goes, you can usually rely on a scientific audience knowing how it works (since it’s covered in most intro stats classes). However, it’s still a good idea to be explicit about stating the null hypothesis (briefly!) because the null hypothesis can be different depending on what you’re using the test for. For instance, in the cards example my null hypothesis was that all the four suit probabilities were identical (i.e., P1=P2=P3=P4=0.25), but there’s nothing special about that hypothesis. I could just as easily have tested the null hypothesis that P1=0.7 and P2=P3=P4=0.1 using a goodness of fit test. So it’s helpful to the reader if you explain to them what your null hypothesis was. Also, notice that I described the null hypothesis in words, not in maths. That’s perfectly acceptable. You can describe it in maths if you like, but since most readers find words easier to read than symbols, most writers tend to describe the null using words if they can.
- A “stat block” is included. When reporting the results of the test itself, I didn’t just say that the result was significant, I included a “stat block” (i.e., the dense mathematical-looking part in the parentheses), which reports all the “raw” statistical data. For the chi-square goodness of fit test, the information that gets reported is the test statistic (that the goodness of fit statistic was 8.44), the information about the distribution used in the test (χ2 with 3 degrees of freedom, which is usually shortened to χ2(3)), and then the information about whether the result was significant (in this case p<.05). The particular information that needs to go into the stat block is different for every test, and so each time I introduce a new test I’ll show you what the stat block should look like.173 However the general principle is that you should always provide enough information so that the reader could check the test results themselves if they really wanted to.
- The results are interpreted. In addition to indicating that the result was significant, I provided an interpretation of the result (i.e., that people didn’t choose randomly). This is also a kindness to the reader, because it tells them something about what they should believe about what’s going on in your data. If you don’t include something like this, it’s really hard for your reader to understand what’s going on.174
As with everything else, your overriding concern should be that you explain things to your reader. Always remember that the point of reporting your results is to communicate to another human being. I cannot tell you just how many times I’ve seen the results section of a report or a thesis or even a scientific article that is just gibberish, because the writer has focused solely on making sure they’ve included all the numbers, and forgotten to actually communicate with the human reader.
comment on statistical notation (advanced)
Satan delights equally in statistics and in quoting scripture
– H.G. Wells
If you’ve been reading very closely, and are as much of a mathematical pedant as I am, there is one thing about the way I wrote up the chi-square test in the last section that might be bugging you a little bit. There’s something that feels a bit wrong with writing “χ2(3)=8.44”, you might be thinking. After all, it’s the goodness of fit statistic that is equal to 8.44, so shouldn’t I have written X2=8.44 or maybe GOF=8.44? This seems to be conflating the sampling distribution (i.e., χ2 with df=3) with the test statistic (i.e., X2). Odds are you figured it was a typo, since χ and X look pretty similar. Oddly, it’s not. Writing χ2(3)=8.44 is essentially a highly condensed way of writing “the sampling distribution of the test statistic is χ2(3), and the value of the test statistic is 8.44”.
In one sense, this is kind of stupid. There are lots of different test statistics out there that turn out to have a chi-square sampling distribution: the X2 statistic that we’ve used for our goodness of fit test is only one of many (albeit one of the most commonly encountered ones). In a sensible, perfectly organised world, we’d always have a separate name for the test statistic and the sampling distribution: that way, the stat block itself would tell you exactly what it was that the researcher had calculated. Sometimes this happens. For instance, the test statistic used in the Pearson goodness of fit test is written X2; but there’s a closely related test known as the G-test175 , in which the test statistic is written as G. As it happens, the Pearson goodness of fit test and the G-test both test the same null hypothesis; and the sampling distribution is exactly the same (i.e., chi-square with k−1 degrees of freedom). If I’d done a G-test for the cards data rather than a goodness of fit test, then I’d have ended up with a test statistic of G=8.65, which is slightly different from the X2=8.44 value that I got earlier; and produces a slightly smaller p-value of p=.034. Suppose that the convention was to report the test statistic, then the sampling distribution, and then the p-value. If that were true, then these two situations would produce different stat blocks: my original result would be written X2=8.44, χ2(3),p=.038, whereas the new version using the G-test would be written as G=8.65, χ2(3), p=.034. However, using the condensed reporting standard, the original result is written χ2(3)=8.44, p=.038, and the new one is written χ2(3)=8.65, p=.034, and so it’s actually unclear which test I actually ran.
So why don’t we live in a world in which the contents of the stat block uniquely specifies what tests were ran? The deep reason is that life is messy. We (as users of statistical tools) want it to be nice and neat and organised… we want it to be designed, as if it were a product. But that’s not how life works: statistics is an intellectual discipline just as much as any other one, and as such it’s a massively distributed, partly-collaborative and partly-competitive project that no-one really understands completely. The things that you and I use as data analysis tools weren’t created by an Act of the Gods of Statistics; they were invented by lots of different people, published as papers in academic journals, implemented, corrected and modified by lots of other people, and then explained to students in textbooks by someone else. As a consequence, there’s a lot of test statistics that don’t even have names; and as a consequence they’re just given the same name as the corresponding sampling distribution. As we’ll see later, any test statistic that follows a χ2 distribution is commonly called a “chi-square statistic”; anything that follows a t-distribution is called a “t-statistic” and so on. But, as the X2 versus G example illustrates, two different things with the same sampling distribution are still, well, different.
As a consequence, it’s sometimes a good idea to be clear about what the actual test was that you ran, especially if you’re doing something unusual. If you just say “chi-square test”, it’s not actually clear what test you’re talking about. Although, since the two most common chi-square tests are the goodness of fit test and the independence test (Section 12.2), most readers with stats training can probably guess. Nevertheless, it’s something to be aware of.


